|
3/13/2024 0 Comments Bcftools filter vcf 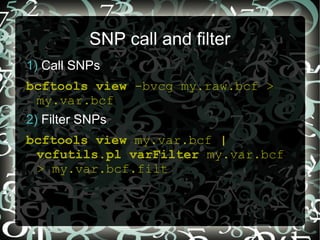
It also includes different scores obtained during sequencing, alignment, and calling to allow quality filtering as well as added sequence annotations to allow annotation-driven filtering. VCF is a tabular text format that provides rich information about each position different from the reference genome. Since the expansion of the 1000 genome project, the Variant Call Format has become more and more popular and is today the default format to represent sequence variation. 5.1 use bcftools_0.2.0 stats and plot-vcfstats_0.2.0 to perform QC on VCF data.4 review bcftools (samtools) variant calls.3.4 call high quality SNP and Indel variants using the Broad GATK. 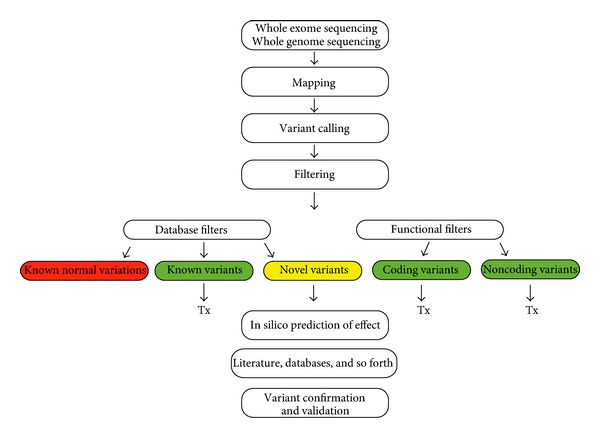
3.3.3 call germline SNVs and Indels separately.3.3.2 call germline SNVs and Indels together.3.3.1.3 method for somatic CNV variants.3.2.2 the script using the recent htslib versions of samtools and bcftools.3.2 call variants with samtools version 1.x and bcftools 1.x (both using htslib).3.1 call variants with samtools and samtools bcftools.2 About bgzip-compressed VCF data and indexing.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
